Dear scientist:
Hi! I am trying to establish a model in some Fluxnet sites to calculate the changes in stomatal conductance of vegetation over the years. I used the fluxnet2015 data to make atmosphere forcing data. I choose compset with 2000_DATM%1PT_CLM50%BGC- CROP_SICE_SOCN_MOSART_SGLC_SWAV. I try to use DATA_MODE =CLM1PT and data_mode=GSWPS3V1.
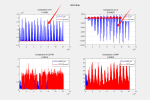
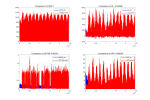
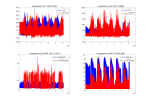
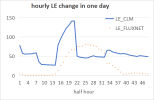
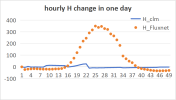
But, When I used DATA_MODE =CLM1PT, the output FSDS had an error different from the input FSDS. Every day FSDS is abnormally high at 17:00-18:00, this leads FSH to an abnormal high in output result.
Then, I tried to change data_mode=GSWPS3V1, FSDS was correct, and the output FSDS was the same as the input FSDS. but it has error Bottom layer specific humidity sent from the atmosphere model is less than zero. I have checked all RH in the input data, It didn't have a value less than zero, I think this may be induced by PSFR and TBOT.
I have mainly three questions:
1, Why does DATA_MODE =CLM1PT make output FSDS become abnormally ? input FSDS is different from putout FSDS. Could you give me some suggestions?
2, when I used data_mode=GSWPS3V1, PSFR(pressure) and TBOT (temperature) lead Bottom layer specific humidity sent from the atmosphere model is less than zero, how to deal with input data to solve the question?
3, compset 2000_DATM%1PT_CLM50%BGC- CROP_SICE_SOCN_MOSART_SGLC_SWAV,Is the compset I have chosen reasonable? Is this compset using the Medlyn method to calculate leaf stomatal conductance? Can I use satellite phenology (I1PtClm50SpGs)?
4,If I choose compset I1PtClm50SpGs,can I use other kind DATA_MODE, such as GSWPS3v1?
The following is my code to run the model:
./create_newcase --case ${SITENAME} --compset 2000_DATM%1PT_CLM50%BGC-CROP_SICE_SOCN_MOSART_SGLC_SWAV --res CLM_USRDAT --mach myintel --compiler intel --run-unsupported
cd ${SITENAME}
./xmlchange DATM_MODE=CLM1PT
./xmlchange CALENDAR=GREGORIAN
./xmlchange DIN_LOC_ROOT_CLMFORC=/home/wjp/cesm/inputdata/fluxnet/
./xmlchange CLM_USRDAT_NAME=${SITENAME}
./xmlchange ATM_DOMAIN_FILE=domain.lnd.1x1pt_${SITENAME}_1x1pt_${SITENAME}.240109.nc
./xmlchange LND_DOMAIN_FILE=domain.lnd.1x1pt_${SITENAME}_1x1pt_${SITENAME}.240109.nc
./xmlchange LND_DOMAIN_PATH=/home/wjp/cesm/inputdata/fluxnet/${SITENAME}/
./xmlchange ATM_DOMAIN_PATH=/home/wjp/cesm/inputdata/fluxnet/${SITENAME}/
./xmlchange RUN_STARTDATE=${yearstart}-01-01
./xmlchange DATM_CLMNCEP_YR_START=${yearstart}
./xmlchange DATM_CLMNCEP_YR_ALIGN=${yearstart}
./xmlchange DATM_CLMNCEP_YR_END=${yearend}
./xmlchange STOP_OPTION=nyears
./xmlchange STOP_N=${yearnum}
./case.setup
echo "fsurdat='/home/wjp/cesm/inputdata/fluxnet/"${SITENAME}"/surfdata_1x1pt_"${SITENAME}"_78pfts_CMIP6_simyr2000_c240109.nc'" >> user_nl_clm
echo "hist_nhtfrq = -1" >> user_nl_clm
echo "hist_mfilt = 8760" >> user_nl_clm
echo "hist_fincl1= 'GPP'" >> user_nl_clm
echo "hist_fincl1='FV'" >> user_nl_clm
./preview_namelists
cp CaseDocs/datm.streams.txt.CLM1PT.CLM_USRDAT user_datm.streams.txt.CLM1PT.CLM_USRDAT
cat CaseDocs/datm_in >> user_nl_datm
./preview_namelists
./check_input_data
./case.build --skip-provenance-check
cd ~/cesm/scratch/${SITENAME}/run
mkdir -p timing/checkpoints
mpirun -np 1 ../bld/cesm.exe
tar -cf ~/${SITENAME}.tar -P $(find . -name ""${SITENAME}".clm2.h0.*.nc")
cd ~
let ist=ist+1
done
Hi! I am trying to establish a model in some Fluxnet sites to calculate the changes in stomatal conductance of vegetation over the years. I used the fluxnet2015 data to make atmosphere forcing data. I choose compset with 2000_DATM%1PT_CLM50%BGC- CROP_SICE_SOCN_MOSART_SGLC_SWAV. I try to use DATA_MODE =CLM1PT and data_mode=GSWPS3V1.
But, When I used DATA_MODE =CLM1PT, the output FSDS had an error different from the input FSDS. Every day FSDS is abnormally high at 17:00-18:00, this leads FSH to an abnormal high in output result.
Then, I tried to change data_mode=GSWPS3V1, FSDS was correct, and the output FSDS was the same as the input FSDS. but it has error Bottom layer specific humidity sent from the atmosphere model is less than zero. I have checked all RH in the input data, It didn't have a value less than zero, I think this may be induced by PSFR and TBOT.
I have mainly three questions:
1, Why does DATA_MODE =CLM1PT make output FSDS become abnormally ? input FSDS is different from putout FSDS. Could you give me some suggestions?
2, when I used data_mode=GSWPS3V1, PSFR(pressure) and TBOT (temperature) lead Bottom layer specific humidity sent from the atmosphere model is less than zero, how to deal with input data to solve the question?
3, compset 2000_DATM%1PT_CLM50%BGC- CROP_SICE_SOCN_MOSART_SGLC_SWAV,Is the compset I have chosen reasonable? Is this compset using the Medlyn method to calculate leaf stomatal conductance? Can I use satellite phenology (I1PtClm50SpGs)?
4,If I choose compset I1PtClm50SpGs,can I use other kind DATA_MODE, such as GSWPS3v1?
The following is my code to run the model:
./create_newcase --case ${SITENAME} --compset 2000_DATM%1PT_CLM50%BGC-CROP_SICE_SOCN_MOSART_SGLC_SWAV --res CLM_USRDAT --mach myintel --compiler intel --run-unsupported
cd ${SITENAME}
./xmlchange DATM_MODE=CLM1PT
./xmlchange CALENDAR=GREGORIAN
./xmlchange DIN_LOC_ROOT_CLMFORC=/home/wjp/cesm/inputdata/fluxnet/
./xmlchange CLM_USRDAT_NAME=${SITENAME}
./xmlchange ATM_DOMAIN_FILE=domain.lnd.1x1pt_${SITENAME}_1x1pt_${SITENAME}.240109.nc
./xmlchange LND_DOMAIN_FILE=domain.lnd.1x1pt_${SITENAME}_1x1pt_${SITENAME}.240109.nc
./xmlchange LND_DOMAIN_PATH=/home/wjp/cesm/inputdata/fluxnet/${SITENAME}/
./xmlchange ATM_DOMAIN_PATH=/home/wjp/cesm/inputdata/fluxnet/${SITENAME}/
./xmlchange RUN_STARTDATE=${yearstart}-01-01
./xmlchange DATM_CLMNCEP_YR_START=${yearstart}
./xmlchange DATM_CLMNCEP_YR_ALIGN=${yearstart}
./xmlchange DATM_CLMNCEP_YR_END=${yearend}
./xmlchange STOP_OPTION=nyears
./xmlchange STOP_N=${yearnum}
./case.setup
echo "fsurdat='/home/wjp/cesm/inputdata/fluxnet/"${SITENAME}"/surfdata_1x1pt_"${SITENAME}"_78pfts_CMIP6_simyr2000_c240109.nc'" >> user_nl_clm
echo "hist_nhtfrq = -1" >> user_nl_clm
echo "hist_mfilt = 8760" >> user_nl_clm
echo "hist_fincl1= 'GPP'" >> user_nl_clm
echo "hist_fincl1='FV'" >> user_nl_clm
./preview_namelists
cp CaseDocs/datm.streams.txt.CLM1PT.CLM_USRDAT user_datm.streams.txt.CLM1PT.CLM_USRDAT
cat CaseDocs/datm_in >> user_nl_datm
./preview_namelists
./check_input_data
./case.build --skip-provenance-check
cd ~/cesm/scratch/${SITENAME}/run
mkdir -p timing/checkpoints
mpirun -np 1 ../bld/cesm.exe
tar -cf ~/${SITENAME}.tar -P $(find . -name ""${SITENAME}".clm2.h0.*.nc")
cd ~
let ist=ist+1
done