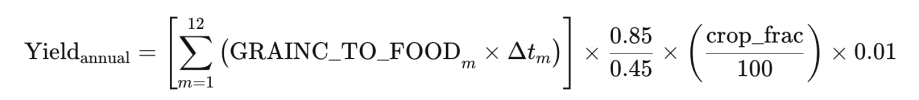What version of the code are you using?
CESM2.2 (CLM5, CAM6-chem/MOSAIC)
Composet: FCnudged
Have you made any changes to files in the source tree?
./xmlchange CLM_BLDNML_OPTS="-bgc bgc -crop" #turn on BGC and crop model
user_nl_cam: Nudge_Model =.false.
user_nl_clm: use_ozone = .true. #Turn on ozone damage
Describe every step you took leading up to the problem:
1. Convert monthly mean flux (GRAINC_TO_FOOD) to annual accumulated grain carbon:
Δtm is seconds in month, and crop_frac represents the fraction (%) of cropland within the grid cell. "0.01" is unit conversion factor from g/m2 to tonnes/ha.
According to the CLM documentation (Section 2.26.2.4.4, Harvest), crop yield is calculated as:

I think the unit for this calculation method is g/m2.
Describe your problem or question:
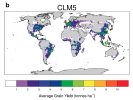
I would appreciate any comments on the discrepancy in my results. Could this be related to an issue in how I calculated crop yield (tonnes/ha)?
CESM2.2 (CLM5, CAM6-chem/MOSAIC)
Composet: FCnudged
Have you made any changes to files in the source tree?
./xmlchange CLM_BLDNML_OPTS="-bgc bgc -crop" #turn on BGC and crop model
user_nl_cam: Nudge_Model =.false.
user_nl_clm: use_ozone = .true. #Turn on ozone damage
Describe every step you took leading up to the problem:
- I output the variable GRAINC_TO_FOOD at monthly frequency
- I then post-process the output to calculate annual crop yield (tonnes/ha)
1. Convert monthly mean flux (GRAINC_TO_FOOD) to annual accumulated grain carbon:
Δtm is seconds in month, and crop_frac represents the fraction (%) of cropland within the grid cell. "0.01" is unit conversion factor from g/m2 to tonnes/ha.
According to the CLM documentation (Section 2.26.2.4.4, Harvest), crop yield is calculated as:

I think the unit for this calculation method is g/m2.
Describe your problem or question:
- Should crop fraction (PCT_LANDUNIT) be applied in yield calculation?
- Or is GRAINC_TO_FOOD already normalized over crop area?
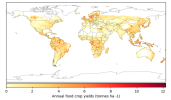
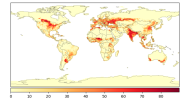
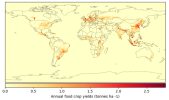
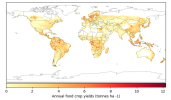
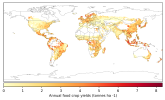
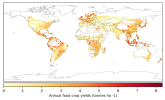
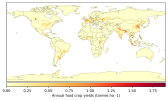
- When I compare my annual crop yield (left panel) with that from Lawrence et al., (2019) (the right panel):


I would appreciate any comments on the discrepancy in my results. Could this be related to an issue in how I calculated crop yield (tonnes/ha)?