Hi everyone,
One of the challenges a lot of new CESM users encounter is that of installing the software to begin with - so we've been working on a 'containerized' CESM environment to help people get up and running, and learning CESM, more easily. Containers offer portable environments - so this works on Macs, Linux and even Windows systems, and runs on anything from personal laptops to high-end servers. Best of all, it requires zero configuration. You can download the image, run it, and have a working CESM environment immediately.
In addition, we've added a Jupyter Lab environment to the image ("CESM-Lab"), giving users a choice of interfaces - shell, or Jupyter notebooks. Since everything is preconfigured, we're also able to provide tutorials on using CESM - the CESM-Lab image comes with a Quick Start notebook walking new users through the main steps of creating a case, configuring it, building it and running it. We plan on adding more tutorials in the future, covering much more, including analysis and visualization of results via Jupyter Notebooks.
A Google Doc with instructions on downloading and installing 'CESM-Lab-2.2' is available here:
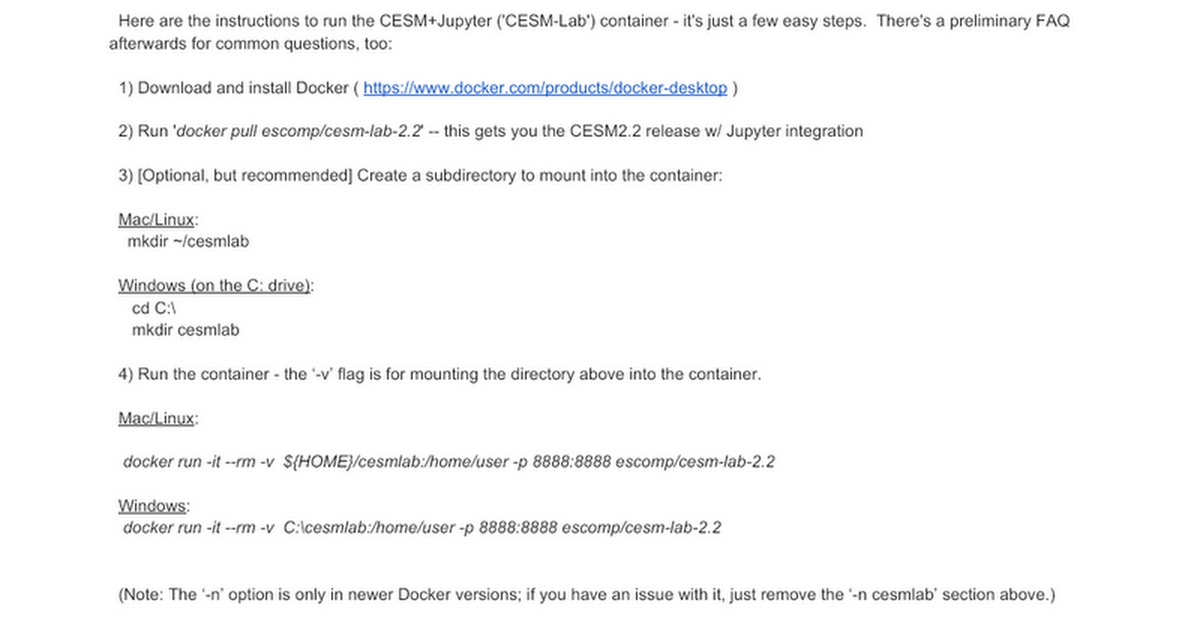
 docs.google.com
docs.google.com
And a spreadsheet of tested compsets, their RAM requirements, input data sizes and performance on both a 4-core laptop and a 36-core server is viewable here. We're adding more to this regularly too:
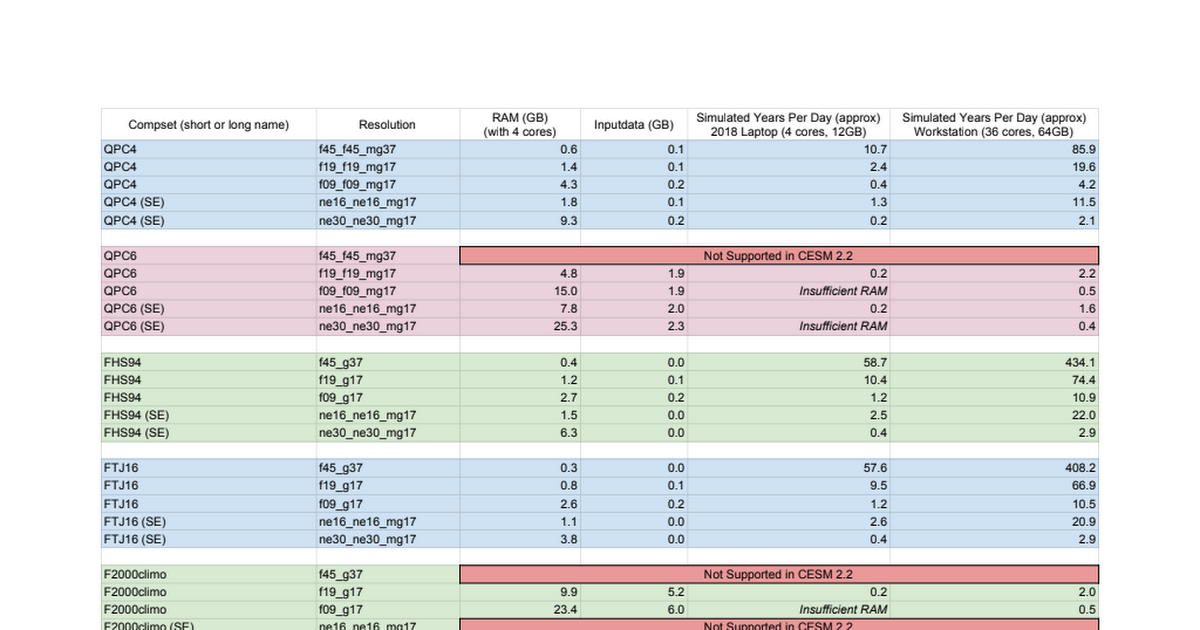
 docs.google.com
docs.google.com
Note that this comes with a full release of CESM 2.2; it's not limited in any way, other than your system resources. RAM limits what compsets and grid sizes you can use, and the type and count of processors will determine the performance of those runs. Some things, like 'simpler model' atmosphere runs or single-point land runs, work fine even with limited resources, whereas fully-coupled 1-degree runs would require a workstation or server with lots of memory.
Finally, this is a technology preview of our container efforts; we're aiming for an official release in the near future. In the meantime, we wanted to get it out to the community, get feedback on how it works for you, and start working on a variety of improvements and additional tutorial material, too!
Thanks, and please ask if you have any questions!
- Brian
One of the challenges a lot of new CESM users encounter is that of installing the software to begin with - so we've been working on a 'containerized' CESM environment to help people get up and running, and learning CESM, more easily. Containers offer portable environments - so this works on Macs, Linux and even Windows systems, and runs on anything from personal laptops to high-end servers. Best of all, it requires zero configuration. You can download the image, run it, and have a working CESM environment immediately.
In addition, we've added a Jupyter Lab environment to the image ("CESM-Lab"), giving users a choice of interfaces - shell, or Jupyter notebooks. Since everything is preconfigured, we're also able to provide tutorials on using CESM - the CESM-Lab image comes with a Quick Start notebook walking new users through the main steps of creating a case, configuring it, building it and running it. We plan on adding more tutorials in the future, covering much more, including analysis and visualization of results via Jupyter Notebooks.
A Google Doc with instructions on downloading and installing 'CESM-Lab-2.2' is available here:
Instructions for using CESM-Lab (Container)
Here are the instructions to run the CESM+Jupyter ('CESM-Lab') container - it's just a few easy steps. There's a preliminary FAQ afterwards for common questions, too: 1) Download and install Docker ( https://www.docker.com/products/docker-desktop ) 2) Run 'docker pull escomp/cesm-lab-2.2...
And a spreadsheet of tested compsets, their RAM requirements, input data sizes and performance on both a 4-core laptop and a 36-core server is viewable here. We're adding more to this regularly too:
Containerized CESM - Tested Configs
Note that this comes with a full release of CESM 2.2; it's not limited in any way, other than your system resources. RAM limits what compsets and grid sizes you can use, and the type and count of processors will determine the performance of those runs. Some things, like 'simpler model' atmosphere runs or single-point land runs, work fine even with limited resources, whereas fully-coupled 1-degree runs would require a workstation or server with lots of memory.
Finally, this is a technology preview of our container efforts; we're aiming for an official release in the near future. In the meantime, we wanted to get it out to the community, get feedback on how it works for you, and start working on a variety of improvements and additional tutorial material, too!
Thanks, and please ask if you have any questions!
- Brian
