Hello CESM community,
I am encountering one ERROR recently when I run a model with the following compset, turning on fire,
compset: I2000Clm50BgcCru
CLM_BLDNML_OPTS="-fire_emis"
After creating the case, I made some changes in user_nl_clm, such as
fire_emis_specifier = 'CB1 = 0.5*BC', 'CB2=0.5*BC', 'BIGALK=BIGALK',.... (its a long list)
I tried running the model by commenting/removing fire_emis_specifier, and the model runs well, which means the problem is within it. I also tried deleting just 'BIGALK=BIGALK' then the error shifted to the next component.
Any suggestions on how can I resolve this problem.
Many thanks!!
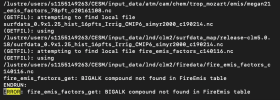
ERROR: fire_emis_factors_get: BIGALK compound not found in FireEmis table
I am encountering one ERROR recently when I run a model with the following compset, turning on fire,
compset: I2000Clm50BgcCru
CLM_BLDNML_OPTS="-fire_emis"
After creating the case, I made some changes in user_nl_clm, such as
fire_emis_specifier = 'CB1 = 0.5*BC', 'CB2=0.5*BC', 'BIGALK=BIGALK',.... (its a long list)
I tried running the model by commenting/removing fire_emis_specifier, and the model runs well, which means the problem is within it. I also tried deleting just 'BIGALK=BIGALK' then the error shifted to the next component.
Any suggestions on how can I resolve this problem.
Many thanks!!
ERROR: fire_emis_factors_get: BIGALK compound not found in FireEmis table